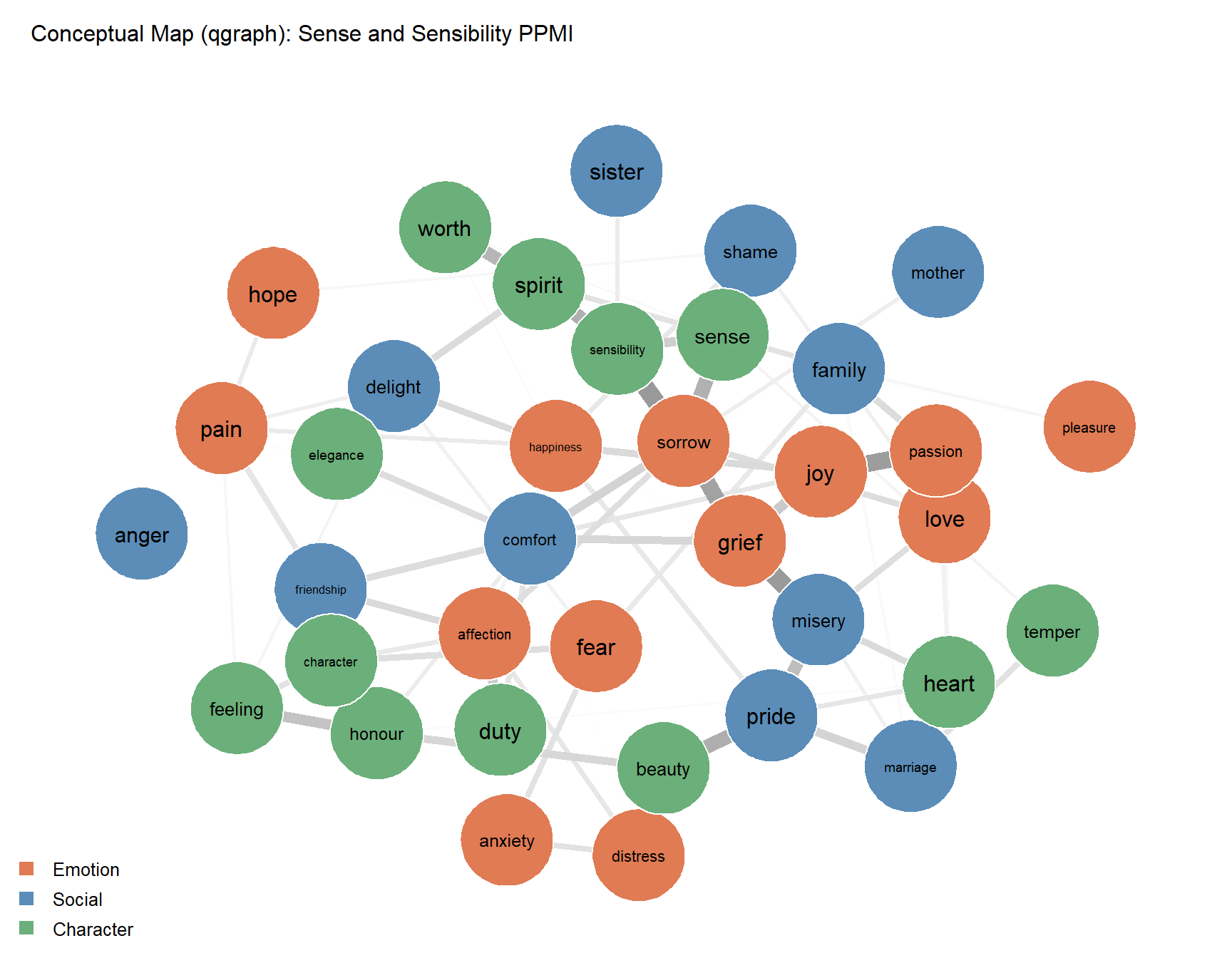
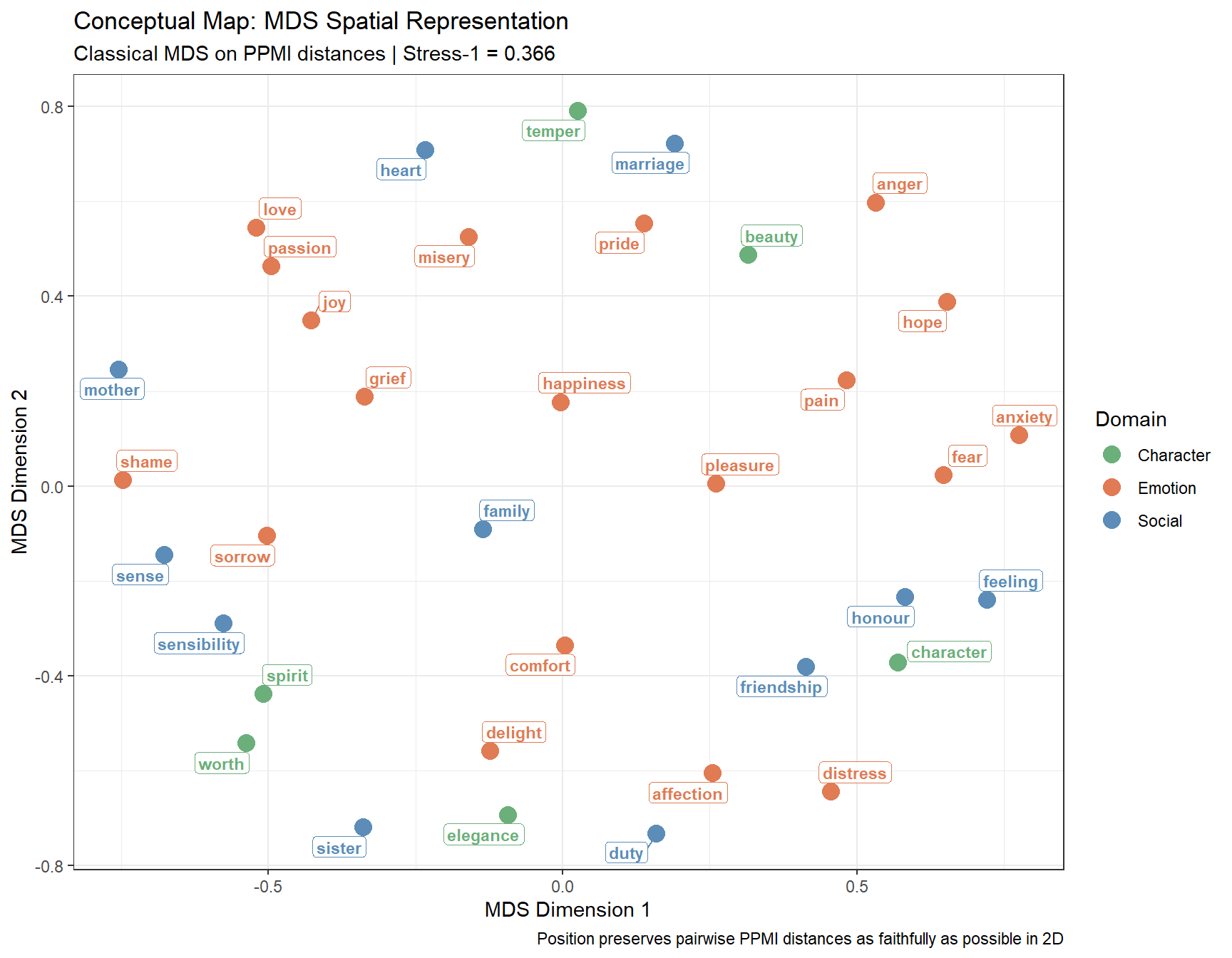
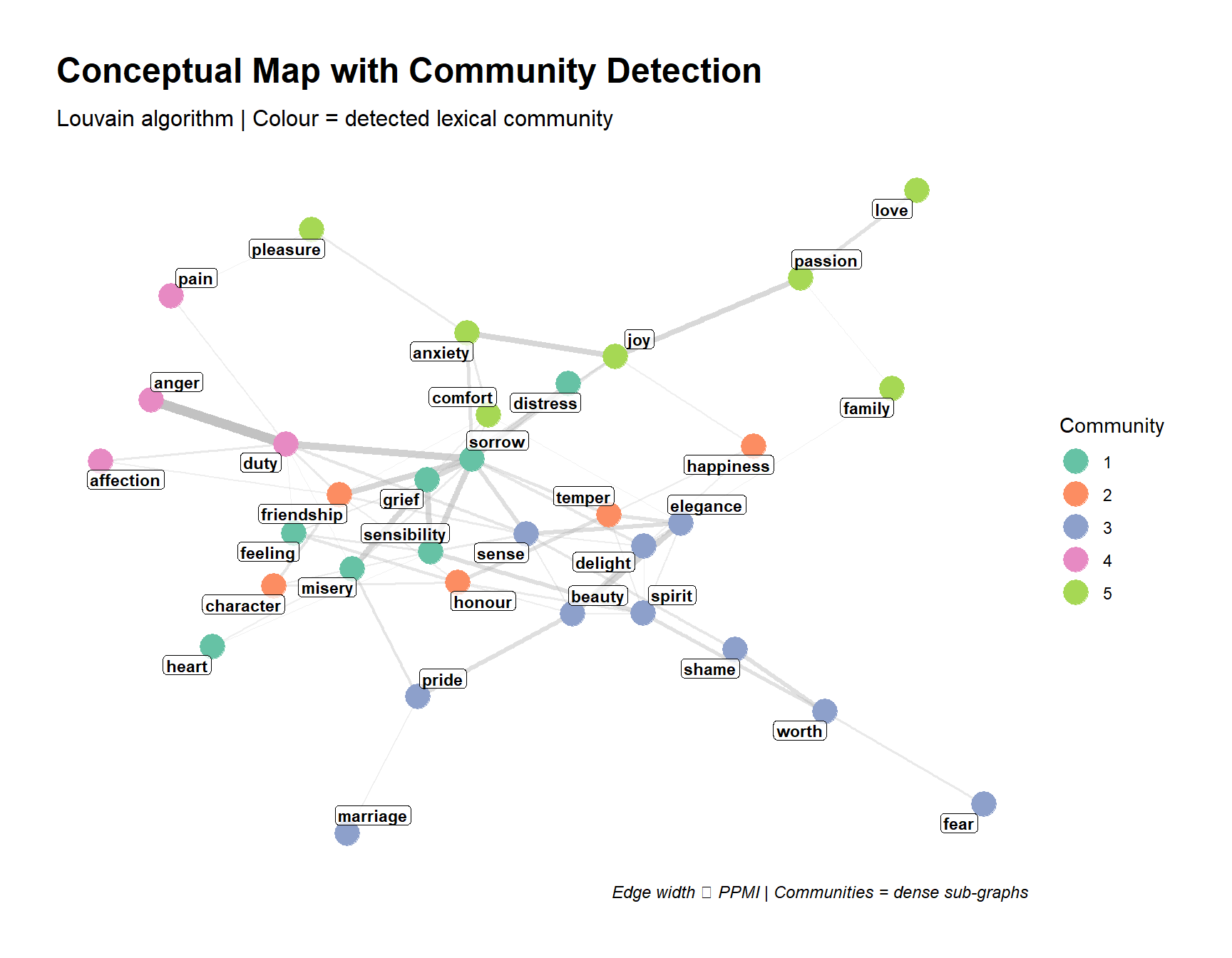
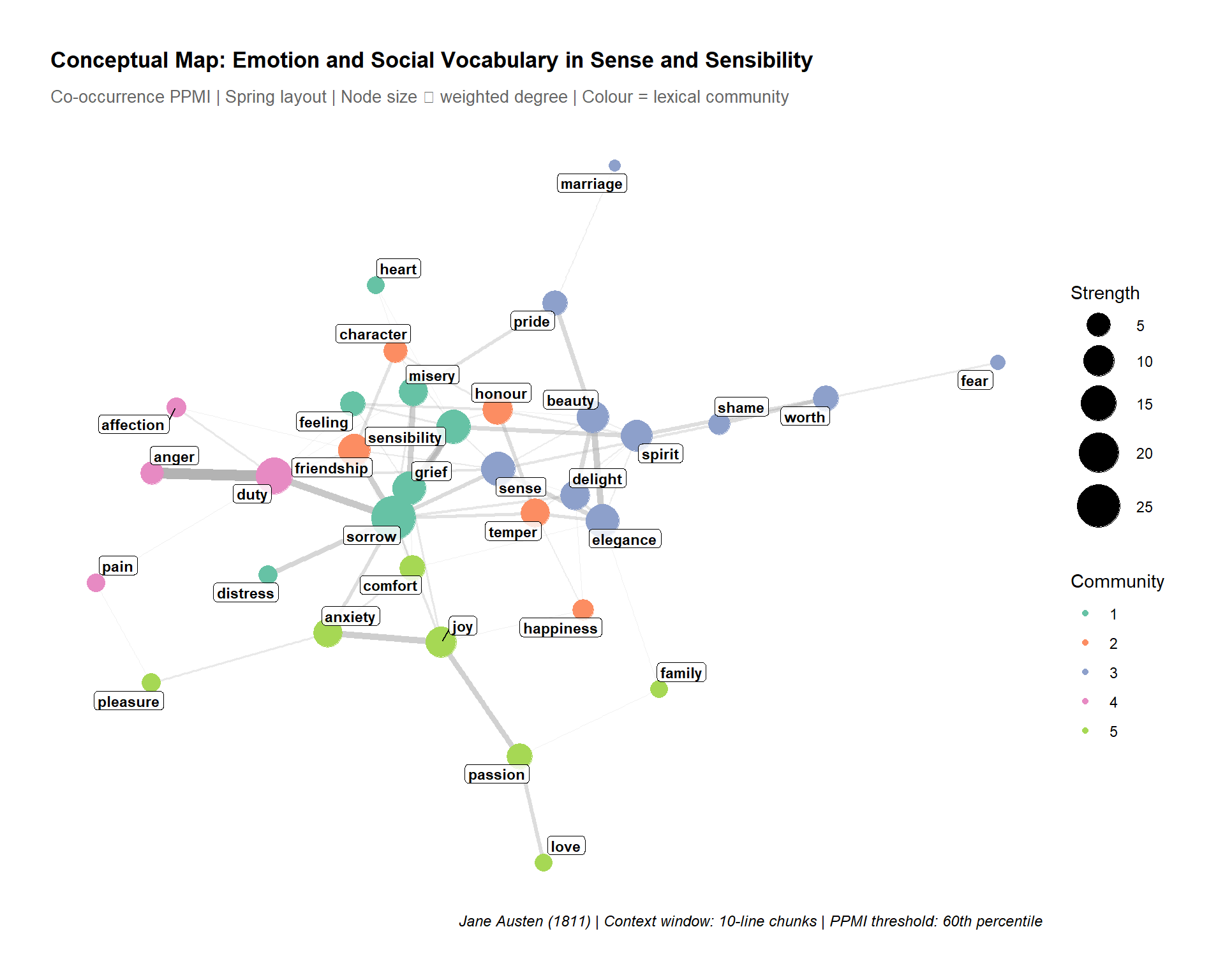
Epskamp, Sacha, Angélique OJ Cramer, Lourens J Waldorp, Verena D Schmittmann, and Denny Borsboom. 2012. “Qgraph: Network Visualizations of Relationships in Psychometric Data.” Journal of Statistical Software 48: 1–18.
Firth, John R. 1957. “A Synopsis of Linguistic Theory, 1930–1955.” In Studies in Linguistic Analysis, 1–32. Oxford: Blackwell.
Fruchterman, Thomas MJ, and Edward M Reingold. 1991. “Graph Drawing by Force-Directed Placement.” Software: Practice and Experience 21 (11): 1129–64.
Harris, Zellig S. 1954. “Distributional Structure.” Word 10 (2-3): 146–62.
Hendrickx, Julien M, and V Blondel. 2008. “Graphs and Networks for the Analysis of Autonomous Agent Systems.” PhD thesis, Catholic University of Louvain, Louvain-la-Neuve, Belgium.
Schneider, Gerold. 2024. “The Visualisation and Evaluation of Semantic and Conceptual Maps.” Linguistics Across Disciplinary Borders: The March of Data. Bloomsbury Publishing (UK), London, 67–94.